NIH-Funded 'scSurvival' AI Pinpoints High-Risk Cancer Cells to Predict Patient Outcomes
OHSU researchers unveil scSurvival, an open-source AI model that uses single-cell RNA sequencing to predict cancer outcomes and identify high-risk cells.
Researchers at Oregon Health & Science University (OHSU) have developed an open-source artificial intelligence model, scSurvival, capable of predicting cancer survival by analyzing tumors at the resolution of individual cells. Funded by the National Institutes of Health (NIH), the study marks a significant shift from traditional diagnostic methods that often rely on averaging genetic data across an entire tissue sample.
Published in the journal Cancer Discovery on April 21, 2026, the research details how scSurvival identifies not only which patients are at high risk but also the specific tumor cells driving that risk. By utilizing single-cell RNA sequencing (scRNA-seq) data, the model can discern subtle cellular differences that traditional "bulk" sequencing typically obscures.
Moving Beyond the Average
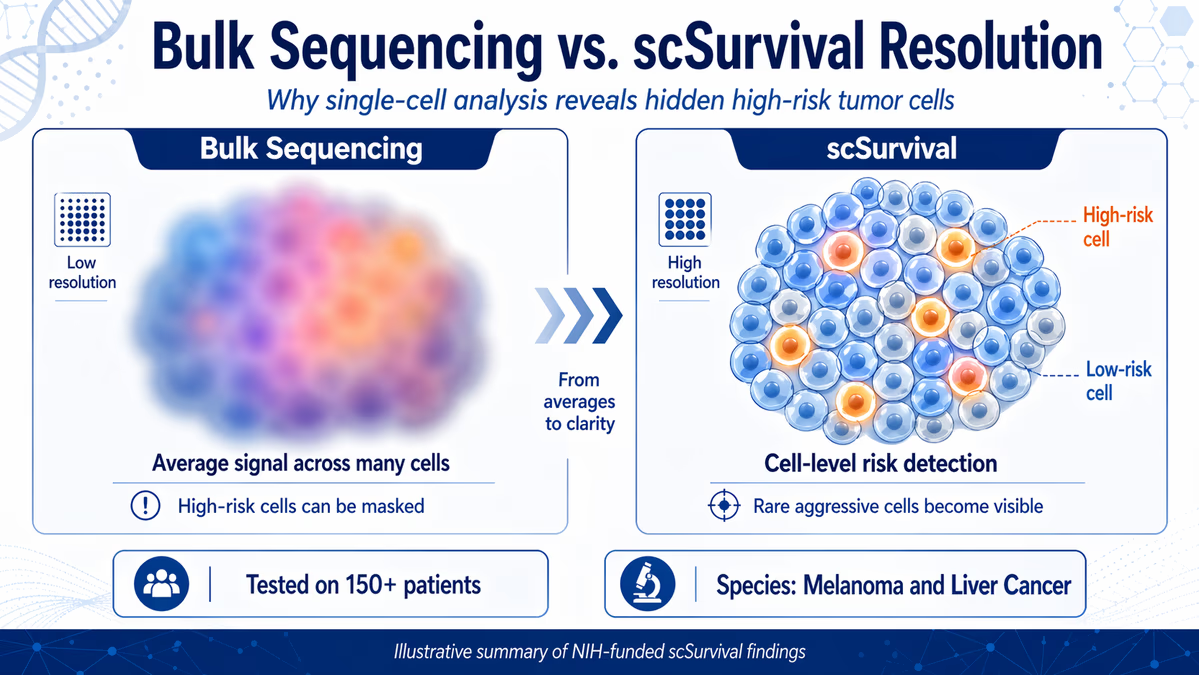
Traditional cancer assessments often employ bulk sequencing, which blends genetic material from millions of cells into a single average. While useful, this approach can hide crucial "heterogeneity"—the variation between cells within a single tumor. Because a small population of resistant or aggressive cells can determine a patient's prognosis, averaging can lead to the loss of vital signals.
"Tumors are very complex, and important signals can be lost when data are averaged across thousands or millions of cells," explains Faming Zhao, Ph.D., a postdoctoral fellow in cancer biology at OHSU and co-lead author of the study. Zhao notes that by examining survival at single-cell resolution, researchers can better understand why patients with the same type of cancer often experience vastly different clinical outcomes.
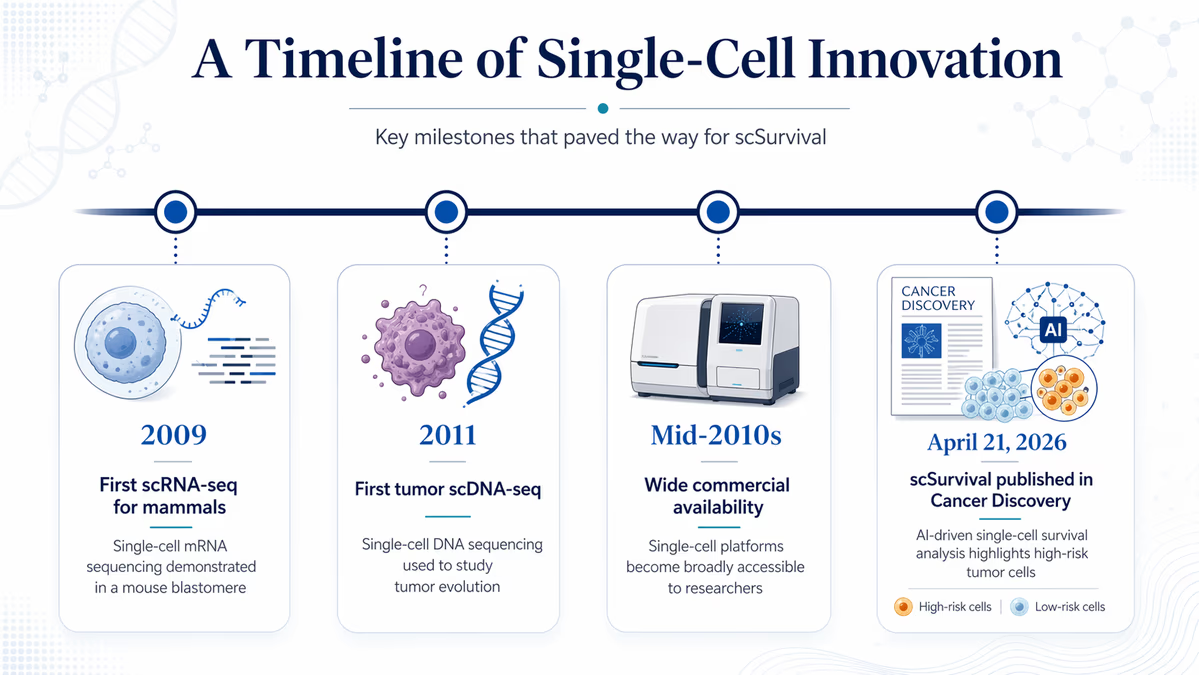
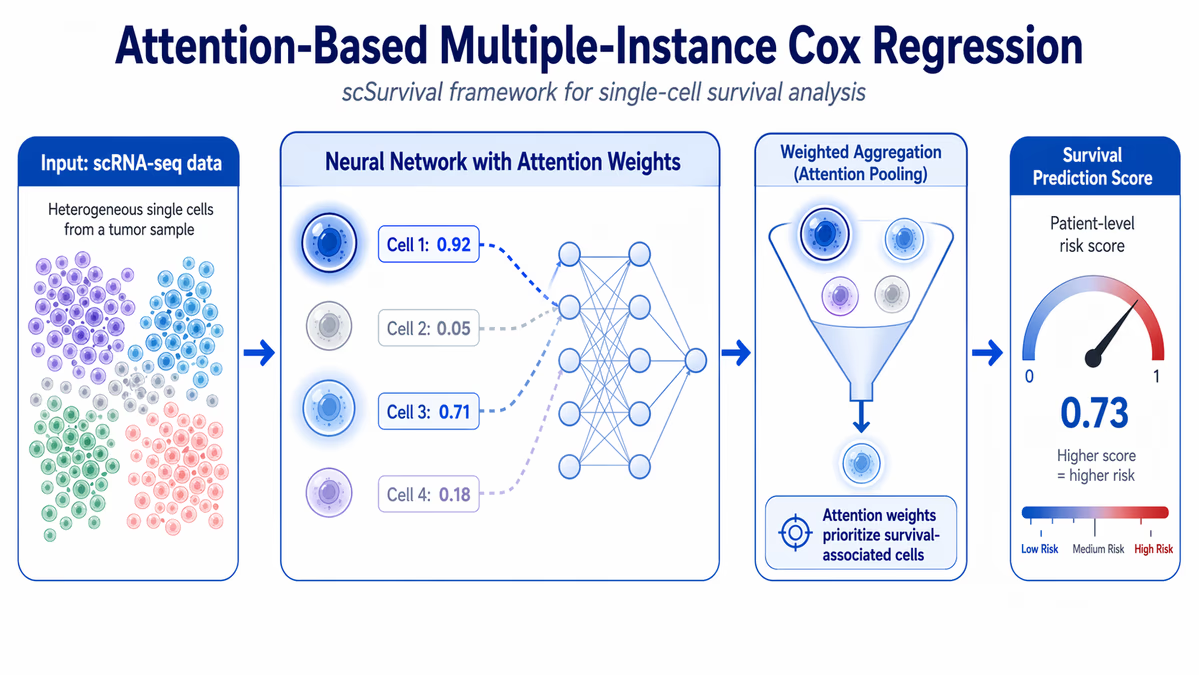
The advent of single-cell sequencing in the late 2000s provided the raw data needed to see these differences, but translating that massive volume of information into reliable survival predictions has remained a hurdle. scSurvival bridges this gap using an advanced machine learning architecture known as an attention-based multiple-instance Cox regression framework.
How scSurvival Works
The core of the scSurvival model is its "attention mechanism." In AI terms, an attention mechanism allows a model to focus on specific parts of a dataset that are most relevant to a specific outcome. In this case, the model assigns a numerical weight to each individual cell based on its predicted influence on patient survival.
Zheng Xia, Ph.D., an associate professor of biomedical engineering at OHSU and the study’s corresponding author, describes the process as taking a "fine-tooth comb" to single-cell data. This allows the model to consider the varying influence that individual cells have on disease progression and survival outcomes. Xia emphasizes that this model is significantly more complex than traditional machine learning approaches, enabling the capture of biological information that was previously inaccessible.
The model was rigorously tested on clinical data from over 150 patients, focusing specifically on melanoma and liver cancer. By analyzing these samples, scSurvival successfully identified specific cell populations—including both immune and tumor cells—that were linked to either favorable or adverse prognoses.
Clinical Implications and Personalized Medicine
The ability to link individual tumor cells directly to patient outcomes is a first for the field. Tao Ren, a postdoctoral fellow in mathematics at OHSU and co-lead author, points out that this is the first kind of single-cell survival analysis that creates this direct connection.
For clinicians, the tool serves a dual purpose. According to Anthony Letai, M.D., Ph.D., director of the NIH's National Cancer Institute (NCI), a risk assessment tool that provides clues as to "why" a patient is at higher risk, rather than just identifying the risk itself, could be transformative for treating difficult cancers. This insight is particularly valuable for identifying patients likely to respond to immunotherapies, where specific immune-cell dynamics play a decisive role.
The researchers have ensured that scSurvival is accessible to the broader scientific community. The program, along with comprehensive tutorials, has been released as open-source software on platforms including GitHub, Zenodo, and Code Ocean. This transparency is expected to accelerate the validation of the model and its integration into various oncology research pipelines worldwide.
The Future of AI in Oncology
The development of scSurvival follows a series of AI advancements in cancer research. From the 2023 UCLA model predicting outcomes via epigenetic factors to the 2025 Stanford "MUSK" model that integrates medical imaging with pathology reports, AI is increasingly becoming the backbone of personalized oncology. The NCI has also signaled its continued support for this field, recently doubling funding for initiatives like the Childhood Cancer Data Initiative to $100 million.
As sequencing costs continue to fall and throughput increases, models like scSurvival are poised to become standard tools in clinical settings. By pinpointing the exact cellular drivers of disease, the medical community moves closer to a future where treatment plans are not just based on the type of cancer a patient has, but on the unique cellular landscape of their specific tumor.